Multiscale Algorithms for Molecular Cimulations
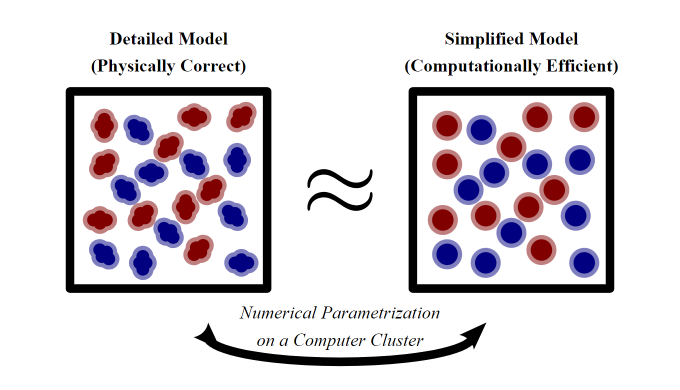
Molecular Cimulation is a powerful tool for probing nanoscale phenomena. Conventional approaches often describe molecules with all their geometrically detailed atoms, but such a computationally inefficient model can solely examine small systems over a short time period. For exhaustively investigating large systems over a long time period, contemporary approaches instead describe molecules by just their isotropically simplified centers, but such a model is physically incorrect in most scenarios. For balancing between computational efficiency and physical correctness, a variety of Multiscale Optimizations have evolved in recent decades: Usually in these algorithms, a simplified model is numerically parameterized via the data of a detailed model. Indeed, in our earlier work, we developed such a computational procedure based on the KullbackLeibler Entropy.
Still, such a numerical algorithm may take many many days on a computer cluster. For overcoming this challenge, in our later work, we developed a hybrid model called ‘Relative Resolution’ (RelRes): If molecules are near to each other, they are characterized by geometrically detailed models, but if molecules are far from each other, they are characterized by isotropically simplified models; via a multipole series that is based on energy conservation, an analytical parametrization inherently emerges from the detailed model to the simplified model, and this can be naturally done on a paper sheet in seconds. Most recently, we implemented RelRes in LAMMPS specifically for the LennardJones potential: We comprehensively showed that RelRes very satisfactorily describes the structural and thermal behavior of several alkane liquids, and it does so with a speedup of an order of magnitude, as compared with its standard counterparts in LAMMPS. We are currently moving in the direction of implementing RelRes for Water, which can be especially complementary for experimental studies of Biomimetic Assemblies that span multiple scales.
- Aviel Chaimovich, Scott Shell. “Relative Entropy as a Universal Metric for Multiscale Errors”, Physical Review E 81 [2010].
- Aviel Chaimovich, Scott Shell. “CoarseGraining Errors and Numerical Optimization using a Relative Entropy Framework”, Journal of Chemical Physics 134 [2011].
- Aviel Chaimovich, Kurt Kremer, Christine Peter. “Relative Resolution: A Multipole Approximation at Appropriate Distances”, Physical Review Research 1 [2019].
- Mark Chaimovich, Aviel Chaimovich. “Relative Resolution: A Computationally Efficient Implementation in LAMMPS”, Journal of Chemical Theory and Computation 17 [2021].
Assembly in Water Solutions
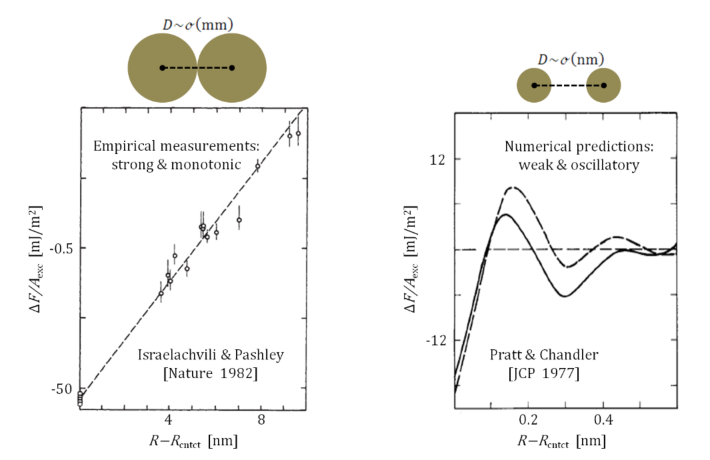
As daVinci expressed it: “Water is the driving Force of all Nature”. Although this is an overstatement, Water is indeed the main constituent of living organisms, and thus, studying Aqueous Solutions is of utmost importance. At the most basic level, scientists study the Hydrophobic Force: How do ‘oily’ (nonpolar) entities aggregate in Water? However, even such a rudimentary question has had a paradox associated with it for almost half of a century: While typical computational predictions suggest a weak oscillatory Force (like the original PrattChandler curve), typical experimental measurements observe a strong monotonic Force (like the original IsraelachviliPashley curve).
Fairly recently, we resolved this paradox for the Hydrophobic Force by performing Molecular Cimulations parametrized via the KullbackLeibler Entropy (in that way, we were capable of probing both small and large systems): Small Hydrophobes aggregate weakly via an oscillatory Force, yet large Hydrophobes aggregate strongly via a monotonic Force; the continuous transition between these two regimes occurs for a Hydrophobe whose size is roughly a nanometer. The sole reason this was not clearly demonstrated before is because the computational predictions and the experimental measurements were being performed on Hydrophobes whose size differs by orders of magnitude! Currently, we are moving in the direction of characterizing the Hydrophilic Force: How do ‘salty’ (polar) entities aggregate in Water? Besides, we are also exploring other aspects of Biomimetic Assembly, specifically those experienced during Force Spectroscopy.
- Aviel Chaimovich, Scott Shell. “LengthScale Crossover of the Hydrophobic Interaction in a CoarseGrained Water Model”, Physical Review E 88 [2013].
- Aviel Chaimovich, Scott Shell. “Tetrahedrality and Structural Order for Hydrophobic Interactions in a CoarseGrained Water Model”, Physical Review E 89 [2014].
